Regulatory Network-centric Dissection of Stress Response
Deciphering molecular stress response is highly relevant understanding how Mtb infects hosts, survive airborne transmission, and adapt to drug treatment. To study how Mtb coordinates stress responses through transcriptional regulation, the Sherman Lab has developed a suite of transcriptional regulatory network-centric tools and characterizations.
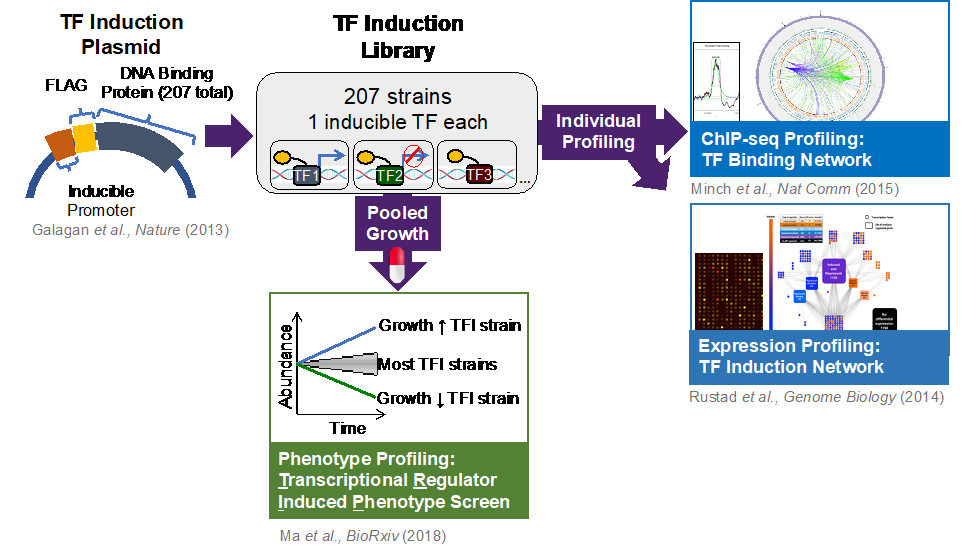
The Sherman Lab has engineered a library of 207 TF-induction (TFI) strains, representing 97% of annotated Mtb regulators. Each strain is engineered for conditional induction of a single Mtb TF and expression of its associated regulon—the set of genes whose expression changes when a given TF is induced[12]. Our ChIP-seq[3] and transcriptome profiling[2] of these TFI strains under in vitro log-phase conditions have revealed a comprehensive baseline network of transcriptional impacts and DNA binding interactions triggered by each TF. With ~16,000 significant binding events1 and ~9000 instances of differential gene regulation[2] this collectively represents the most comprehensive transcriptional regulatory network for any organism.
The Sherman Lab’s novel Transcriptional Regulator Induced Phenotype (TRIP) screen simultaneously quantifies condition-dependent fitness (i.e., growth and survival) phenotypes elicited by every Mtb transcription factor (TF) within a single experiment[4].
The Sherman Lab’s novel Transcriptional Regulator Induced Phenotype (TRIP) screen simultaneously quantifies condition-dependent fitness (i.e., growth and survival) phenotypes elicited by every Mtb transcription factor (TF) within a single experiment[4].
TRIP screens offer several powerful advantages:
1) Emergent phenotypes are accessible, since regulons generally include multiple genes selected for co-regulation by evolution.
2) TF expression is chemically triggered, enabling context-specific interrogation of perturbations.
3) Essential regulators and effector genes can be assessed.
4) TFI fitness phenotypes can be deconstructed to identify candidate effector target genes using our transcriptional regulatory network.
2) TF expression is chemically triggered, enabling context-specific interrogation of perturbations.
3) Essential regulators and effector genes can be assessed.
4) TFI fitness phenotypes can be deconstructed to identify candidate effector target genes using our transcriptional regulatory network.
TRIP is broadly extensible—we have applied the pipeline to study Mtb fitness under numerous stress conditions, including host infection and drug treatment. The fundamental strategy can also be adapted to investigate other species